My goal in this post is to give an overview of Friedman’s Test and then offer R code to perform post hoc analysis on Friedman’s Test results. (The R function can be downloaded from here)
Preface: What is Friedman’s Test
Friedman test is a non-parametric randomized block analysis of variance. Which is to say it is a non-parametric version of a one way ANOVA with repeated measures. That means that while a simple ANOVA test requires the assumptions of a normal distribution and equal variances (of the residuals), the Friedman test is free from those restriction. The price of this parametric freedom is the loss of power (of Friedman’s test compared to the parametric ANOVa versions).
The hypotheses for the comparison across repeated measures are:
- H0: The distributions (whatever they are) are the same across repeated measures
- H1: The distributions across repeated measures are different
The test statistic for the Friedman’s test is a Chi-square with [(number of repeated measures)-1] degrees of freedom. A detailed explanation of the method for computing the Friedman test is available on Wikipedia.
Performing Friedman’s Test in R is very simple, and is by using the “friedman.test” command.
Post hoc analysis for the Friedman’s Test
Assuming you performed Friedman’s Test and found a significant P value, that means that some of the groups in your data have different distribution from one another, but you don’t (yet) know which. Therefor, our next step will be to try and find out which pairs of our groups are significantly different then each other. But when we have N groups, checking all of their pairs will be to perform [n over 2] comparisons, thus the need to correct for multiple comparisons arise.
The tasks:
Our first task will be to perform a post hoc analysis of our results (using R) – in the hope of finding out which of our groups are responsible that we found that the null hypothesis was rejected. While in the simple case of ANOVA, an R command is readily available (“TukeyHSD”), in the case of friedman’s test (until now) the code to perform the post hoc test was not as easily accessible.
Our second task will be to visualize our results. While in the case of simple ANOVA, a boxplot of each group is sufficient, in the case of a repeated measures – a boxplot approach will be misleading to the viewer. Instead, we will offer two plots: one of parallel coordinates, and the other will be boxplots of the differences between all pairs of groups (in this respect, the post hoc analysis can be thought of as performing paired wilcox.test with correction for multiplicity).
R code for Post hoc analysis for the Friedman’s Test
The analysis will be performed using the function (I wrote) called “friedman.test.with.post.hoc”, based on the packages “coin” and “multcomp”. Just a few words about it’s arguments:
- formu – is a formula object of the shape: Y ~ X | block (where Y is the ordered (numeric) responce, X is a group indicator (factor), and block is the block (or subject) indicator (factor)
- data – is a data frame with columns of Y, X and block (the names could be different, of course, as long as the formula given in “formu” represent that)
- All the other parameters are to allow or suppress plotting of the results.
friedman.test.with.post.hoc <- function(formu, data, to.print.friedman = T, to.post.hoc.if.signif = T, to.plot.parallel = T, to.plot.boxplot = T, signif.P = .05, color.blocks.in.cor.plot = T, jitter.Y.in.cor.plot =F)
{
# formu is a formula of the shape: Y ~ X | block
# data is a long data.frame with three columns: [[ Y (numeric), X (factor), block (factor) ]]
# Note: This function doesn't handle NA's! In case of NA in Y in one of the blocks, then that entire block should be removed.
# Loading needed packages
if(!require(coin))
{
print("You are missing the package 'coin', we will now try to install it...")
install.packages("coin")
library(coin)
}
if(!require(multcomp))
{
print("You are missing the package 'multcomp', we will now try to install it...")
install.packages("multcomp")
library(multcomp)
}
if(!require(colorspace))
{
print("You are missing the package 'colorspace', we will now try to install it...")
install.packages("colorspace")
library(colorspace)
}
# get the names out of the formula
formu.names <- all.vars(formu)
Y.name <- formu.names[1]
X.name <- formu.names[2]
block.name <- formu.names[3]
if(dim(data)[2] >3) data <- data[,c(Y.name,X.name,block.name)] # In case we have a "data" data frame with more then the three columns we need. This code will clean it from them...
# Note: the function doesn't handle NA's. In case of NA in one of the block T outcomes, that entire block should be removed.
# stopping in case there is NA in the Y vector
if(sum(is.na(data[,Y.name])) > 0) stop("Function stopped: This function doesn't handle NA's. In case of NA in Y in one of the blocks, then that entire block should be removed.")
# make sure that the number of factors goes with the actual values present in the data:
data[,X.name ] <- factor(data[,X.name ])
data[,block.name ] <- factor(data[,block.name ])
number.of.X.levels <- length(levels(data[,X.name ]))
if(number.of.X.levels == 2) { warning(paste("'",X.name,"'", "has only two levels. Consider using paired wilcox.test instead of friedman test"))}
# making the object that will hold the friedman test and the other.
the.sym.test <- symmetry_test(formu, data = data, ### all pairwise comparisons
teststat = "max",
xtrafo = function(Y.data) { trafo( Y.data, factor_trafo = function(x) { model.matrix(~ x - 1) %*% t(contrMat(table(x), "Tukey")) } ) },
ytrafo = function(Y.data){ trafo(Y.data, numeric_trafo = rank, block = data[,block.name] ) }
)
# if(to.print.friedman) { print(the.sym.test) }
if(to.post.hoc.if.signif)
{
if(pvalue(the.sym.test) < signif.P)
{
# the post hoc test
The.post.hoc.P.values <- pvalue(the.sym.test, method = "single-step") # this is the post hoc of the friedman test
# plotting
if(to.plot.parallel & to.plot.boxplot) par(mfrow = c(1,2)) # if we are plotting two plots, let's make sure we'll be able to see both
if(to.plot.parallel)
{
X.names <- levels(data[, X.name])
X.for.plot <- seq_along(X.names)
plot.xlim <- c(.7 , length(X.for.plot)+.3) # adding some spacing from both sides of the plot
if(color.blocks.in.cor.plot)
{
blocks.col <- rainbow_hcl(length(levels(data[,block.name])))
} else {
blocks.col <- 1 # black
}
data2 <- data
if(jitter.Y.in.cor.plot) {
data2[,Y.name] <- jitter(data2[,Y.name])
par.cor.plot.text <- "Parallel coordinates plot (with Jitter)"
} else {
par.cor.plot.text <- "Parallel coordinates plot"
}
# adding a Parallel coordinates plot
matplot(as.matrix(reshape(data2, idvar=X.name, timevar=block.name,
direction="wide")[,-1]) ,
type = "l", lty = 1, axes = FALSE, ylab = Y.name,
xlim = plot.xlim,
col = blocks.col,
main = par.cor.plot.text)
axis(1, at = X.for.plot , labels = X.names) # plot X axis
axis(2) # plot Y axis
points(tapply(data[,Y.name], data[,X.name], median) ~ X.for.plot, col = "red",pch = 4, cex = 2, lwd = 5)
}
if(to.plot.boxplot)
{
# first we create a function to create a new Y, by substracting different combinations of X levels from each other.
subtract.a.from.b <- function(a.b , the.data)
{
the.data[,a.b[2]] - the.data[,a.b[1]]
}
temp.wide <- reshape(data, idvar=X.name, timevar=block.name,
direction="wide") #[,-1]
wide.data <- as.matrix(t(temp.wide[,-1]))
colnames(wide.data) <- temp.wide[,1]
Y.b.minus.a.combos <- apply(with(data,combn(levels(data[,X.name]), 2)), 2, subtract.a.from.b, the.data =wide.data)
names.b.minus.a.combos <- apply(with(data,combn(levels(data[,X.name]), 2)), 2, function(a.b) {paste(a.b[2],a.b[1],sep=" - ")})
the.ylim <- range(Y.b.minus.a.combos)
the.ylim[2] <- the.ylim[2] + max(sd(Y.b.minus.a.combos)) # adding some space for the labels
is.signif.color <- ifelse(The.post.hoc.P.values < .05 , "green", "grey")
boxplot(Y.b.minus.a.combos,
names = names.b.minus.a.combos ,
col = is.signif.color,
main = "Boxplots (of the differences)",
ylim = the.ylim
)
legend("topright", legend = paste(names.b.minus.a.combos, rep(" ; PostHoc P.value:", number.of.X.levels),round(The.post.hoc.P.values,5)) , fill = is.signif.color )
abline(h = 0, col = "blue")
}
list.to.return <- list(Friedman.Test = the.sym.test, PostHoc.Test = The.post.hoc.P.values)
if(to.print.friedman) {print(list.to.return)}
return(list.to.return)
} else {
print("The results where not significant, There is no need for a post hoc test")
return(the.sym.test)
}
}
# Original credit (for linking online, to the package that performs the post hoc test) goes to "David Winsemius", see:
# http://tolstoy.newcastle.edu.au/R/e8/help/09/10/1416.html
}
Example
(The code for the example is given at the end of the post)
Let's make up a little story: let's say we have three types of wine (A, B and C), and we would like to know which one is the best one (in a scale of 1 to 7). We asked 22 friends to taste each of the three wines (in a blind fold fashion), and then to give a grade of 1 till 7 (for example sake, let's say we asked them to rate the wines 5 times each, and then averaged their results to give a number for a persons preference for each wine. This number which is now an average of several numbers, will not necessarily be an integer).
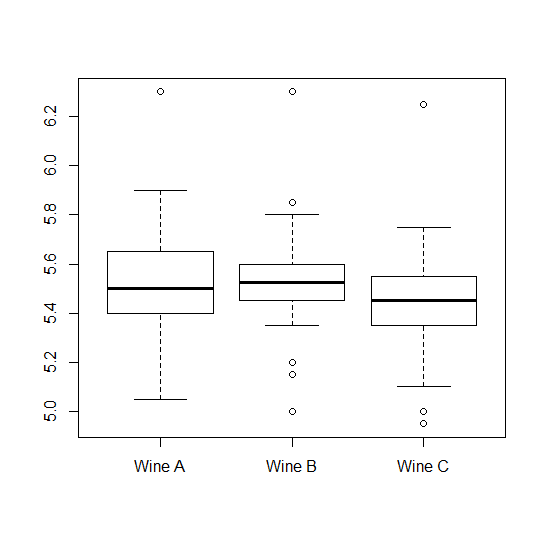
After getting the results, we started by performing a simple boxplot of the ratings each wine got. Here it is:
The plot shows us two things: 1) that the assumption of equal variances here might not hold. 2) That if we are to ignore the "within subjects" data that we have, we have no chance of finding any difference between the wines.
So we move to using the function "friedman.test.with.post.hoc" on our data, and we get the following output:
$Friedman.TestAsymptotic General Independence Testdata: Taste byWine (Wine A, Wine B, Wine C)stratified by TastermaxT = 3.2404, p-value = 0.003421$PostHoc.TestWine B - Wine A 0.623935139Wine C - Wine A 0.003325929Wine C - Wine B 0.053772757
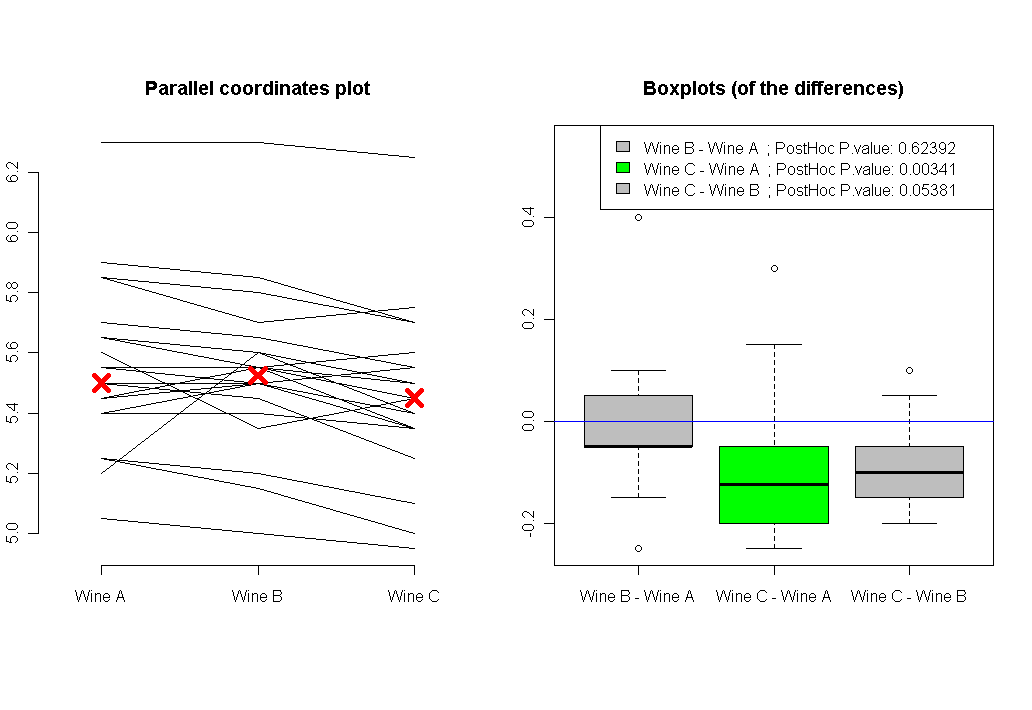
The conclusion is that once we take into account the within subject variable, we discover that there is a significant difference between our three wines (significant P value of about 0.0034). And the posthoc analysis shows us that the difference is due to the difference in tastes between Wine C and Wine A (P value 0.003). and maybe also with the difference between Wine C and Wine B (the P value is 0.053, which is just borderline significant).
Plotting our analysis will also show us the direction of the results, and the connected answers of each of our friends answers:
Here is the code for the example:
source("https://www.r-statistics.com/wp-content/uploads/2010/02/Friedman-Test-with-Post-Hoc.r.txt") # loading the friedman.test.with.post.hoc function from the internet
### Comparison of three Wine ("Wine A", "Wine B", and
### "Wine C") for rounding first base.
WineTasting <- data.frame(
Taste = c(5.40, 5.50, 5.55,
5.85, 5.70, 5.75,
5.20, 5.60, 5.50,
5.55, 5.50, 5.40,
5.90, 5.85, 5.70,
5.45, 5.55, 5.60,
5.40, 5.40, 5.35,
5.45, 5.50, 5.35,
5.25, 5.15, 5.00,
5.85, 5.80, 5.70,
5.25, 5.20, 5.10,
5.65, 5.55, 5.45,
5.60, 5.35, 5.45,
5.05, 5.00, 4.95,
5.50, 5.50, 5.40,
5.45, 5.55, 5.50,
5.55, 5.55, 5.35,
5.45, 5.50, 5.55,
5.50, 5.45, 5.25,
5.65, 5.60, 5.40,
5.70, 5.65, 5.55,
6.30, 6.30, 6.25),
Wine = factor(rep(c("Wine A", "Wine B", "Wine C"), 22)),
Taster = factor(rep(1:22, rep(3, 22))))
with(WineTasting , boxplot( Taste ~ Wine )) # boxploting
friedman.test.with.post.hoc(Taste ~ Wine | Taster ,WineTasting) # the same with our function. With post hoc, and cool plots
If you find this code useful, please let me know (in the comments) so I will know there is a point in publishing more such code snippets...
Hi Tal, very interesting post 🙂 and definitely worthwhile that you continue posting such snippets.
I am a psychology student currently working on the stats for my MA thesis. I have been using repeated measure ANOVAs to analyze pain perception data. However, some of the normality assumptions are very violated (e.g., extreme skew and kurtosis, many outliers).
I have been thinking of investigating a non-parametric approach to this analysis, and your post has piqued my interest in Friedman’s test.
However, is it possible (excuse my ignorance if this is a stupid question ;P) to incorporate between group analyses in a Friedman’s test? For example, my analysis needs to look like this: 2 (between-group 1:a, b) by 2 (between-group 2: a, b) by 6 (within-group 1: a, b, c, d, e, f)
Further, can Friedman’s test be extended to multivariate repeated measure designs? For example, my potential analyses looks like: 2 (between-group 1:a, b) by 2 (between-group 2: a, b) by 3 (within-group 1: a, b, c) by 6 (within-group 2: a, b, c, d, e, f)
And finally, would your post hoc test be able to extend to these more complex designs? or are there other strategies you would recommend?
Anyways, thanks for the post, it was very interesting 🙂
Cheers,
Pat
Hi Pat,
Thank you for your encouraging comment 🙂
To your question: The friedman test is about the “between” group (while considering the “within” group). However, it is only one-way (repeated measures) ANOVA. That means that the more sophisticated models you asked about will not be friedman’s test any more.
What you are asking is if it is possible to do multiway non-parametric anova (with/without repeated measures) and with balanced/unbalanced designs – and such that allows you to do posthoc analysis (and if the tests can also be powerful – it would be great). The bad news is that while it might be possible, I don’t have the experience to tell you how.
The good news though, is that there is a generalization of Wilcoxon-Kruskal-Wallis test that can handle interactions in the model. you are worth having a look at the lrm function in the Design package or the polr function in the MASS package. For a tutorial on ordinal logistic regression in ecology (using lrm), see
@Article{gui00ord,
author = {Guisan, Antoine and Harrell, Frank E.},
title = {Ordinal response regression models in
ecology},
journal = {Journal of Vegetation Science},
year = 2000,
volume = 11,
pages = {617-626},
annote = {teaching;ordinal logistic model}
}
(This answer is thanks to the kind answer of Frank Harell, please see here: https://stat.ethz.ch/pipermail/r-help/2008-January/152333.html)
If you get to explore this better and get to find good tutorials/books/insights – please feel welcome to drop by and let me know about them.
All the best,
Tal
Thanks for your helpful reply Tal, I really appreciate your feedback.
I’ll spend some time investigating the the directions you have pointed me and post back any useful new information that I find 🙂
Cheers and thanks again Tal,
Pat
———————
note: I had a bit of trouble clicking through that link, but was eventually able to find the post at:
https://mailman.stat.ethz.ch/pipermail/r-help/2008-January/152333.html
Sir, what is the difference between Fishers LSD and Friedman’s Test, How to do Fishers LSD and graph it, thanks for the code, with regards, Samuel, Bangalore India
Hello Dear Duleep,
Regarding Fisher LSD:
Fisher LSD is a post hoc test for ANOVA (which is a parametric test).
Friedman’s test is a type of ANOVA (which is nonparametric one way repeated measures).
The code I presented in this post gives both how to do Friedman’s test AND how to do a post hoc analysis on it.
Doing Fishers LSD means you want to do a post hoc analysis, which relies on a t-test. In a quick search I didn’t find a code for doing it. If you come a cross it, please feel welcome to come back and share the link/code.
All the best,
Tal
Update: I got an e-mail from Duleep Samuel, with an answer to his question.
One can do Fishers LSD with the function “LSD.test” (from the “agricolae” package).
The answer to Samuel came from: Felipe author of Agricolae, Rob author of R in action and Todos Logos.
Hi, Hal.
A very usefull snippet indeed. Particularly liked the boxplot of the differences.
Well done.
Thank you for your very useful code. I was wondering though, (I’m a molecular biologist not a statistician) what is the name of this post hoc analysis? Is it Dunn’s multiple comparison test?
Hi Eric,
The post-hoc tests are:
Wilcoxon-Nemenyi-McDonald-Thompson test
Hollander & Wolfe (1999), page 295
The book in question is
Nonparametric Statistical Methods, 2nd Edition [Hardcover]
By
Myles Hollander (Author), Douglas A. Wolfe (Author)
More on it can be viewed here:
http://www.amazon.com/Nonparametric-Statistical-Methods-Myles-Hollander/dp/0471190454
Best,
Tal
Hi Hal,
Like the other posters, i have adopted your code for my own post hoc tests, and ran into a slight bug?
“Error in .local(.Object, …) : ‘x’ is not a balanced factor”
not quite sure to what this is referring… I can send you sample data that produces this error…. do you know what “x” is referring to?
never mind…. missing a data point!
Hello Tal/Joshua. I am getting the same error, but all of my rows are complete. Did you mean one row was incomplete when you wrote “missing a data point”?
Why does this error occur?
I would be awesome if you could help me out.
Was an answer to this ever found?
I would like to know as well,
Hi Tal,
Sorry for this question, but I’m not sure of what is the “block” and what is the “group” in my case. Would you be ok to help me?
I have 5 individuals, and a measure for each individual 10 times (“sessions”). I would like to see the effect of sessions apart from effect of individuals.
Anyway thank you for your helpful post.
Sab
Hi Sab,
Thanks for visiting 🙂
Your “block” is the 5 individuals, and the “group” is the “sessions”.
Although notice that when doing this for 10 sessions, the resulting plots will be VERY big (all the 10 over 2 permutations = 45 plots).
Consider doing something like merging the different sessions, and checking it on only (for example) quarters of sessions. (If you first looked at the data on how many mergers to make, know that the validity of your choice was a bit shaky, see: http://en.wikipedia.org/wiki/Data_dredging)
Hi, I wonder if it is correct to use the Wilcoxon as a posthoc for Friedman and Mann-Whitney as a posthoc for Kruskall-Wallis?
Tank you!
Anna
Hi Annabel,
That is not what I used in this post, but I believe you can do that. However, you would need to adjust the P values you get (for example, by using the Bonferroni correction).
The advantage of this post hoc method is that it is (probably) more powerful then what you will get using Bonferroni (which can be too conservative sometimes)
Hi Tal and Annabel,
As a late comment on this aspect of the discussion, I’d say that the following should also be a good (and simple) post-hoc test :
pairwise.wilcox.test(x=data$measure,g=data$group,p.adjust.method=”holm”,paired=T)
In this example there are several measures in each group (one measure per block). Here, the block is implicit (it is taken into account through paired=T). The Holm method for multiple-testing corrections is less conservative than Bonferroni.
Best wishes,
Gael
Hi Tal,
I was wondering if it’s possible to do a power analysis for Friedman test (i.e., computing the power).
thanks,
Peggy
Hi Peggy.
I don’t know of any existing script that does that. But you could write such a script using bootstaping.
Although you would need to define the structure of your data and then implement the simulation. Not too complex, but takes time and careful thinking.
If you would write such a think, please come back to share.
If you wish to have more help on this, I suggest sending a massage to one of the R mailing lists to see if someone can maybe point to a solution.
Best,
Tal
Hi.
Regarding question from Pat:
If you have ordinal data from say a likert scale, and you got two groups, how do you analyse these data? In SPSS i cannot see any option to analyse for interaction effects between two groups in Friedmans test. It is all good when you have only 1 group, but with 2 groups it seems as if Friedmans test is useless.
Regards
Hello Jacob,
You are absolutely correct.
The Friedman test is only of use when we are doing a one-way repeated measures (non parametric) ANOVA.
I was wondering myself how this can be performed when the need arise for a multiway ANOVA. As I wrote on my other post repeated measures anova with r (tutorials and functions), I couldn’t (yet) find a solution for this issue. (although there is a solution for multi way, non-parametric, NOT repeated measures, ANOVA. See that post for more on that).
If you ever come about a solution for this case, please let me know.
Best,
Tal
Hi!
Wow, this place has been a huge help to me, thank you, Tal.
I’ve gotten a bit confused though, in my search for the correct statistics for my master thesis in behavioral biology.
I am looking at 10 trainingsessions (over time) with a number of stress related symptoms within each session. These symptoms are a result of a human impact on 10 different animals.
Using the Friedman test I’ve already established that there is a significant difference in the number of symptoms between the 10 sessions. Now I would like to know between which sessions the difference is. Have I understood correctly if I think I can test this with the method mentioned above (the post hoc Friedman)?
Furthermore I would like to make a trendline from session 1 to session 10 showing if the number of symptoms are in- or declining over time. Do you know how to do this in R?
Hope all this makes sence…
Best wishes,
Katrine
Hi Katrine,
I am glad you found my post helpful 🙂
If I understood you correctly, your dataset is of 10 animals. For each animal you’ve got 10 (overtime) observation. Each observation is a number (the number of stress factors).
If I got you correctly, then indeed the code in this post (for the post hoc friedman test) could potentially help you.
BUT, I don’t think it will be very helpful.
You are dealing with a case of 10 repeated measures on each individual animal. That means you’ve got 10 over 2 comparisons (45). That is a big number.
Also, you are having a time effect here, why not use it in some way ?
For example, why not only look at 3 time points (start, middle, finish) and make the post hoc comparison on them?
This post hoc might offer more easily interpretable results.
A better yet solution would be to go further and see if you can use some mixed models on your data.
Then, what you are looking to check is if you are having a significant slope for the trend line.
Although I haven’t yet wrote about these methods (and I also don’t know how your data is behaving and how legitimate it is to use mixed models on it).
Consider also giving a look to what I wrote here:
https://www.r-statistics.com/2010/04/repeated-measures-anova-with-r-tutorials/
But if my earlier tip (of just using less data points – to allow interpretation), is not enough for you – then I suggest you try and find some professional help to look at your data and see if mixed models can work on it or not. (Or just learn it by yourself, but it might take some time 🙂 )
Regarding the visualization, it is very straight forward.
This tutorial:
http://www.ats.ucla.edu/stat/R/seminars/Repeated_Measures/repeated_measures.htm
Shows how to do it with lattice. You might also want to try ggplot2 for that (both have some learning curve, but offer good results).
Good luck,
Tal
Hi Tal,
Thanks a lot for all your helpfull information.
I have used your code for R to perform a post hoc in a Friedman test. I would like to know how can I cite your code in a journal.
Thanks,
Luis
Hello LuisM,
Thank you very much for offering me the honor of being cited.
Due to my lack of experience, I might be missing on how this should be done, but here is how you might do it:
The analysis was done using R:
@Manual{,
title = {R: A Language and Environment for Statistical
Computing},
author = {{R Development Core Team}},
organization = {R Foundation for Statistical Computing},
address = {Vienna, Austria},
year = 2010,
note = {{ISBN} 3-900051-07-0},
url = {http://www.R-project.org}
}
With the “coin” and “multcomp” packages.
Performing the post-hoc tests of:
Wilcoxon-Nemenyi-McDonald-Thompson test
Hollander & Wolfe (1999), page 295
Using the code of “Tal Galili”, published on r-statistics.com (https://www.r-statistics.com/2010/02/post-hoc-analysis-for-friedmans-test-r-code)
Thanks Tal!!
Hi Tal,
I can’t wait to successfully use the code you have worked out and so kindly shared with us ALL.
To start. i’ve plugged in the following
friedman.test.with.post.hoc <- function(formu, data, to.print.friedman = T, to.post.hoc.if.signif = T, to.plot.parallel = T, to.plot.boxplot = T, signif.P = .05, color.blocks.in.cor.plot = T, jitter.Y.in.cor.plot =F)
I receive a + symbol. Is this correct? I've been working at trying to get my data into your code for 2 days, but am new to R and am struggling.
I have updated the coin, multcomp, and colorspace with success. Does this need to be done in a specific order or each time I reopen R?
I have 3 columns of data. Id = turtle name (block), temperature (factor)…collected stomach temperature from 7 turtles) and time (group)…want to see if there is a difference in stomach temperature for 4 time periods (dawn, day, dusk, night). I do not have 0s or blanks in my data set and found a sig. difference for Friedman.test in R.
any advice would be greatly appreciated!!!
Awesome work. It’s really helping me. I add your blog in my bookmark !
Hi,
Great that such a script is available, really appreciate it.
Doesn’t seem to work on all data though. For e.g., I have ordinal data ranging from 1-3 in groups = 15, and blocks = 7.
R produces this error:
Error in data[, X.name] : incorrect number of dimensions
The ordinary friedman.test gives me:
Friedman chi-squared = 50.127, df = 14, p-value = 5.814e-06
So in principle your script should also work. But I am at a loss to explain why it does not. Will keep trying though.
Thanks again!
To those that read my comment, please just ignore it: the error was entirely down to my own lack of experience with R and attempting to name a variable ‘data’. The script works perfectly!
What modifications I need to the r code for reading SAS datasets? Would I need an SAS ODBC driver or SAS/Share?
Thanks
Bharat
Hello Bharat,
I never tried importing SAS files, but you can look here:
https://cran.r-project.org/doc/manuals/R-data.html#EpiInfo-Minitab-SAS-S_002dPLUS-SPSS-Stata-Systat
Cheers,
Tal
Hi Tal
Congratulation for your blog, it is very helpful and at very good level. My questions is due my lack of knowledge in statistic, and unfortunately not related with R. Altough, I will appreciate you advices. Can I perform a repeated measures ANOVA on ranks (Friedman’s test), if i have some missing values (different n values) in some of the treatments?
Thanks
As far as I know, such observations will be omitted.
Hi tal
Its very nice information.
Can you analyse my data for Posthoc LSD? what is the meaning of letters a, ab, or ac on chart? on which basis this letters are mentioned on the chart?
Hello
Purushottam ,
The letters simply indicate to you which of the groups are found distinct from one another.
So if you have the groups one, two , three. And they are found in post hoc to be:
a one
ab two
b three
That means that one is different from three, but that we can’ distinguish between one and two, or two from three.
I hope this helps.
Best,
Tal
Hi Tal,
excellent blog/post.
Nice design, good writing style, elucidating content, generous code sharing and respectful citation of sources… that’s how it should be. Congratulations.
I am looking for a non-parametric two-way test and I thought Friedman would do the job, while you are stating that it is a one-way test.
It looks like your answer to Pat seems to point in the right direction. I will give it a try.
I do really wonder why there is no friedman.test() in vanilla R.
Tanks a lot and keep up the good work,
Reinhard
Oops, there is a
friedman.test()
See
?friedman.test
🙂
Did first think of it while writing the post before …
Hi there
Question about your test statistic. I’m getting really different p-values running a Friedman’s Test in SAS vs. the friedman.test function in R. I know that SAS is using the somewhat updated (as of Iman and Davenport 1980) version of the test statistic, as presented in 2nd and 3rd editions of Conover. The R help does not cite a formula, but does cite an older text (1973). So I’m wondering if the function in R is using the old test statistic? You appear to use a newer version of the same Hollander and Wolfe text, but when I use your code I get a p-value more similar to that provided by the friedman.test function. But not exactly the same. Any insight on (a) what test statistic is being used in R, and why your test and the friedman.test function are not identical for the global test? Thanks very much!
R
Okay, so cancel the bit about SAS vs. R. I’m getting the same results now. But I would still love to know why your p-values are slightly different from the friedman.test function (what is the maxT method?), and especially why I’m seeing two very slightly different p-values associated with the same maxT statistic (one before and one after the multiple comparisons). Anyway. Sorry for the previous post, which was mostly wrong!
You use Tukey in your code-isn’t this usually reserved for parametric analysis?
Thanks
No.
‘Tukey’ simply corresponds to the particular set of contrasts that represents all pairwise comparisons of the groups. So, for three groups (A, B and C) you’d want the contrasts representing B vs A, C vs A and C vs B.
This was really helpful! thanks
My pleasure 🙂
Hi~
Your code is so charming!but I got some trouble with it…I have an data frame like this S T TD
1 0.00 1 D1
2 0.00 1 D2
3 0.00 1 D3
4 0.70 1 D4
5 0.74 1 D5
6 0.63 1 D6
7 0.61 1 D7
8 0.56 1 D8
9 0.54 1 D9
10 0.60 1 D10
11 0.00 2 D1
12 0.68 2 D2
13 0.57 2 D3
14 0.56 2 D4
15 0.56 2 D5
16 0.48 2 D6
17 0.57 2 D7
18 0.58 2 D8
19 0.56 2 D9
20 0.56 2 D10
21 0.00 3 D1
22 0.76 3 D2
23 0.68 3 D3
24 0.70 3 D4
25 0.59 3 D5
26 0.57 3 D6
27 0.62 3 D7
28 0.50 3 D8
29 0.59 3 D9
30 0.00 3 D10
31 0.00 4 D1
32 0.76 4 D2
33 0.70 4 D3
34 0.73 4 D4
35 0.68 4 D5
36 0.66 4 D6
37 0.80 4 D7
38 0.55 4 D8
39 0.53 4 D9
40 0.52 4 D10
41 0.00 5 D1
42 0.82 5 D2
43 0.67 5 D3
44 0.50 5 D4
45 0.67 5 D5
46 0.58 5 D6
47 0.00 5 D7
48 0.00 5 D8
49 0.00 5 D9
50 0.00 5 D10
51 0.00 6 D1
52 0.00 6 D2
53 0.59 6 D3
54 0.48 6 D4
55 0.68 6 D5
56 0.62 6 D6
57 0.48 6 D7
58 0.49 6 D8
59 0.48 6 D9
60 0.42 6 D10
61 0.00 7 D1
62 0.00 7 D2
63 0.69 7 D3
64 0.00 7 D4
65 0.00 7 D5
66 0.00 7 D6
67 0.00 7 D7
68 0.57 7 D8
69 0.49 7 D9
70 0.61 7 D10
71 0.00 8 D1
72 0.72 8 D2
73 0.75 8 D3
74 0.00 8 D4
75 0.00 8 D5
76 0.00 8 D6
77 0.00 8 D7
78 0.66 8 D8
79 0.74 8 D9
80 0.56 8 D10and I trend to compute ” friedman.test(S ~ TD | T) “but I can’t run your procedure successfully…it just gave me a wrong message :”In if (to.post.hoc.if.signif) { :
the condition has length > 1 and only the first element will be used”how can I do next ?
Nice coding & example!
Hi Jan, thank you – I am glad it was helpful for you.
Nice coding & example!
Hi Jan, thank you – I am glad it was helpful for you.
Thanks for this. However, I’m concerned that when I run the same code 10 times on the same data set, I come up with 10 different results (they’re not very different, but it concerns me, and leaves me in a quandary when it comes to reporting a value).
Please excuse my ignorance, but could you please explain where the random factor comes in?
Hi Ben,
The
friedman.test.with.post.hoc
Function is based on the:
symmetry_test
function.
In which the calculation of the distribution appears to include some error in it. The option I use in this function is of distribution = c(“asymptotic”) . When reading asymptotic function help file, it reads that it relies on the
abseps
parameter, (wich is “double, the absolute error tolerance”)
So you could probably increase the precision of the function.
And make sure to use some set.seed before running it so your results will be reproducible.
I hope this helps.
Tal
Thanks, that helps a lot!
Thank you! This really helped me.
This was very helpful. Thanks so much!
First of all thanks for the script. It’s been quite useful to me.
I would like to comment to issues:
1) I’m working with some data now (19 groups and 8 blocks) and the Friedman rank sum test and the Asymptotic General Independence Test of your script give quite different results (p-values of 0.0002091 and 0.05858, respectively).
I know that this is no so unusual, given that each test it is based on different procedures, but it’s quite puzzling for me and I’m not sure if I should try to another procedure to test differences between groups (e.g. pairwise comparisons followed by Bonferroni correction). I would thank your opinion.
2) With mi current version of R I have the following warnings when I apply your script to data with significant differences:
Warning message:
sd() is deprecated.
Use apply(*, 2, sd) instead.
Thanks
Hello, thank you for the code. It is very helpful, but I have a question regarding the conclusion… What is the conclusion? Would you say that Wine A and Wine B are the best, and Wine C is the worse (even though there is no significant difference between Wine B and Wine C? Or, would you say that there is no difference between the wines?
Thanks for this code!
Great code, it is exactly what I needed. If I use it for comparing data for publication, how should I cite you? Thank you very much
Hello dear Daniel,
Thank you for the kind words and offer.
Most of the code was done by people like the R core team (for R), Torsten Hothorn, Kurt Hornik, Mark A. van de Wiel, Achim Zeileis (for {coin}) and Torsten Hothorn, Frank Bretz and Peter Westfall (for {multcomp})
You can use the citation information for the main code using the following code:
citation()
citation(“coin”)
citation(“multcomp”)
Performing the post-hoc tests of:
Wilcoxon-Nemenyi-McDonald-Thompson test
Hollander & Wolfe (1999), page 295
using the R code of ‘Tal Galili’ (from https://www.r-statistics.com/2010/02/post-hoc-analysis-for-friedmans-test-r-code)
Hiya,
Can I follow up the same unresolved error posted 7 months ago please. I too am getting the error message:
Warning message:
sd() is deprecated.
Use apply(*, 2, sd) instead.
I would be grateful for anyones advice how to resolve this.
Thanks
Hello,
I analyzed my data using this function and it gave me the following error message:
‘Error in mvt(lower = lower, upper = upper, df = 0, corr = corr, delta = mean, :
only dimensions 1 <= n <= 1000 allowed'
Does this mean there is a limit to the amount of data that could be analyzed?
Thank you,
Miguel
Hey there, using friedman.test I get p= .0501, but using friedman.test.with.post.hoc i get p=.0425… Obviously you’d need the data and rest of the details for a full explanation, but is anyone else getting different results?
I’m also getting such differences, even when using the data provided in the example (0.00336 with the code found here, 0.003805 with friedman.test).
With my data, it’s even worse: 0.02498 with code from here, 0.01312 with friedman.test (that’s a very big difference)!
Thanks for sharing this!
Now, suppose that I have a multivariate case where I measure 300 variables for each subject. How should I proceed?
Hi,
Thanks you for your work, I’m really novice in R, but your code seems to be the best for my data analysis. Unfortunatly, I have an error in the beginning of the function, so I can’t manage to use it.
My error is:
Error in if (jitter.Y.in.cor.plot) {:
the argument can not be interpreted as a logical value
I have copy and paste your code, I tried to figure what is wrong, but I can’t find why I have the error. So I can’t even run the function.
Thanks you
Delphine
Thank you for the code. Two quick questions: 1) results are printed twice, almost identical (except a small difference in p); 2) is it applicable also to replicated data? I see warnings on ReshapeWide… thanks.
I have tried stepping through the code, and cannot get one line to work, and am wondering what I am doing wrong. Immediately at line one:
friedman.test.with.post.hoc <- function(Max.Sept~Treatment|Block, data = np_d, to.print.friedman = T, to.post.hoc.if.signif = T, to.plot.parallel = T, to.plot.boxplot = T, signif.P = .05, color.blocks.in.cor.plot = T, jitter.Y.in.cor.plot =F)
I get Error: unexpected '~' in "friedman.test.with.post.hoc <- function(Max.Sept~"
I have also tried installing colorspace and it just restarts R over and over, but never successfully loads.
Hi Tal, Thanks for the helpful code. It is a great function to use. Once we understand that there is a difference between the tastes, is there a statistical way of ranking the Wines, best to worst?
Hi,
I’m having some trouble with this code and couldn’t find anything at google about this error, can you help me?
The error is at line 66 (print(the.sym.test) ). The variable exists, and RStudio says that is a “Formal class MaxTypeIndependenceTest” type.
The error that I got is:
Error in checkmvArgs(lower = lower, upper = upper, mean = mean, corr = corr, :
‘lower’ not specified or contains NA
There is no NA in my data. Do you have a clue of where should I start digging to find the issue?
Thank you!
PS: Btw: thank you for the code!
Hi tal,
The post is very interesting and useful. I was wondering how do I print the rank(mean rank values) for each group to actual show which is the better one.
Thank you!
Respected readers, Can I use the Friedman Test to determine the significance differences of means within sessions in a group?
Hi Tal, thanks for the code!
I used it to analyze data my Masters Thesis in behavioral ecophysiology and will probably also publish it (with reference to you, of course).
I’m intrigued by the Wilcoxon-Nemenyi-McDonald-Thompson test. I haven’t read anything about this test being used before and I can’t find a lot of infos on it. I’m in Germany and can’t get the book by Hollander et al either. Also, I can’t find where you implemented it in your code.The only package I could find to use this test was the NSM3 package that you don’t use in the code.
Do you know, how this test adjusts p-values and what’s the difference to a basic wilcoxon-test? (I already found out that the basic wilcoxon-test has problems with ties and zeroes and therefore wouldn’t work for my data anyways)
Sorry for the questions, I hope you can help me a bit with it.
Cheers and thanks again!
Hello and thanks for your detailed example that help me quite a lot (as a novice psychology student).
Coming to report correctly the output in APA format, I was wondering if the maxT value correspond to the khi2 value ?
As with your example, can we report like this :
“A non parametric Friedman test lead to significant differences between wines (khi2(2, N=22) = 3.2404, p = 0.0034)”
Thanks,
–Frédéric
Hi Tal,
I know its been a while since you made this post, but it has been a lifesaver. My one issue is that I keep getting the following error:
“Error in .local(.Object, …) : ‘x’ is not a balanced factor”
I’m not sure what this is referring to,or what I can do to fix it! Any help would be appreciated! I can also send you my data set.
Thank you!
Hi Tal, thank you for this work. You helped to present data analyses part in my paper so that journal was happy. This post is something that one can not easily find in text books while is absolutely necessary.