This post on the dendextend package is based on my recent paper from the journal bioinformatics (a link to a stable DOI). The paper was published just last week, and since it is released as CC-BY, I am permitted (and delighted) to republish it here in full:
abstract
Summary: dendextend is an R package for creating and comparing visually appealing tree diagrams. dendextend provides utility functions for manipulating dendrogram objects (their color, shape, and content) as well as several advanced methods for comparing trees to one another (both statistically and visually). As such, dendextend offers a flexible framework for enhancing R’s rich ecosystem of packages for performing hierarchical clustering of items.
Availability: The dendextend R package (including detailed introductory vignettes) is available under the GPL-2 Open Source license and is freely available to download from CRAN at: (https://cran.r-project.org/package=dendextend)
Contact: [email protected]
Introduction
Hierarchical Cluster Analysis (HCA) is a widely used family of unsupervised statistical methods for classifying a set of items into some hierarchy of clusters (groups) according to the similarities among the items. The R language (R Core Team, 2014) – a leading, cross-platform, and open source statistical programming environment – has many implementations of HCA algorithms (Hornik, 2014; Chipman and Tibshirani, 2006; Witten and Tibshirani, 2010; Schmidtlein et al., 2010). The output of these various algorithms are stored in the hclust object class, while the dendrogram class is an alternative object class that is often used as the go-to intermediate representation step for visualizing an HCA output.
In many R packages, a figure output is adjusted by supplying the plot function with both an object to be plotted and various graphical parameters to be modified (colors, sizes, etc.). However, different behavior happens in the (base R) plot.dendrogram function, in which the function is given a dendrogram object that contains within itself (most of) the graphical parameters to be used when plotting the tree. Internally, the dendrogram class is represented as a nested list of lists with attributes for colors, height, etc. (with useful methods from the stats package). Until now, no comprehensive framework has been available in R for flexibly controlling the various attributes in dendrogram’s class objects.
The dendextend package aims to fill this gap by providing a significant number of new functions for controlling a dendrogram’s structure and graphical attributes. It also implements methods for visually and statistically comparing different dendrogram objects. The package is extensively validated through unit-testing (Wickham, 2011), offers a C++ speed-up (Eddelbuettel and François, 2011) for some of the core functions through the dendextendRcpp package, and includes three detailed vignettes.
The dendextend package is primarily geared towards HCA. For phylogeny analysis, the phylo object class (from the ape package) is recommended (Paradis et al., 2004). A comprehensive comparison of dendextend, ape, as well as other software for tree analysis, is available in the supplementary materials.
Description
Updating a dendrogram for visualization
The function set(dend, what, value), in dendextend, accepts a dendrogram (i.e.: dend) as input and returns it after some adjustment. The parameter what is a character indicating the property of the tree to be adjusted (see Table 1) based on value. The user can repeatedly funnel a tree, through different configuration of the set function, until a desired outcome is reached.
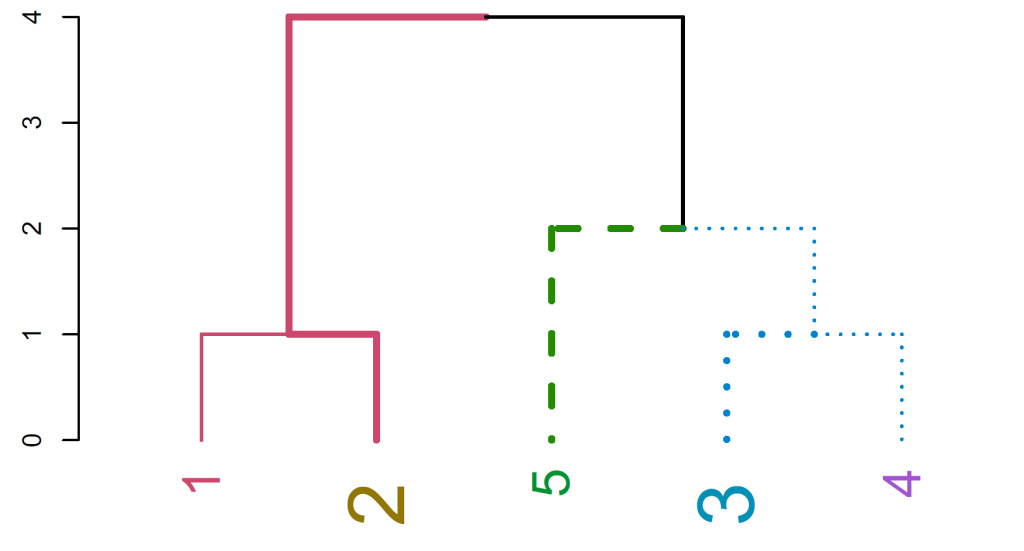
Fig. 1. A dendrogram after modifying various graphical attributes
Fig. 1 is created by clustering a vector of 1 to 5 into a dendrogram:
dend0 <- 1:5 %>% dist %>% hclust %>% as.dendrogram
The above code uses the convenient forward-pipe operator %>% (Milton and Wickham, 2014), which is just like running:
dend0 <- as.dendrogram(hclust(dist(1:5)))
Next, the tree is plotted after repeatedly using the set function:
dend0 %>% set("labels_color") %>%
set("labels_cex", c(2,3)) %>%
set("branches_lwd", c(2,4)) %>% set("branches_k_lty", k=3) %>% set("branches_k_color", k = 3)%>% plotThe “value” vector is recycled in a depth-first fashion, with the root node considered as having a branch (which is not plotted by default). The parameters of the new tree can be explored using the functions get_nodes_attr and get_leaves_attr. Also, we can rotate and prune a tree with the respective functions.
Table 1. Available options for the “what” parameter when using the set function for adjusting the look of a dendrogram
| Description | Option name |
| Set the labels’ names, color (per color, or with k clusters), size, turn to character | labels, labels_to_character, labels_colors, labels_cex, labels_to_character |
| Set the leaves’ point type, color, size, height | leaves_pch, leaves_col, leaves_cex, hang_leaves |
| Set all nodes’ point type, color, size | nodes_pch, nodes_col, nodes_cex |
| Set branches’ line type, color, width – per branch, based on clustering the labels, and for specific labels | branches_lty, branches_col, branches_lwd, branches_k_color, by_labels_branches_lty, by_labels_branches_col, by_labels_branches_lwd |
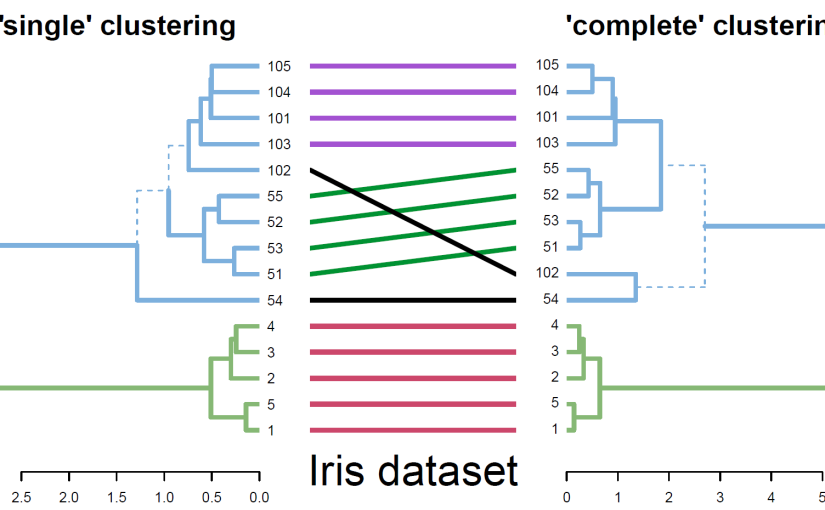
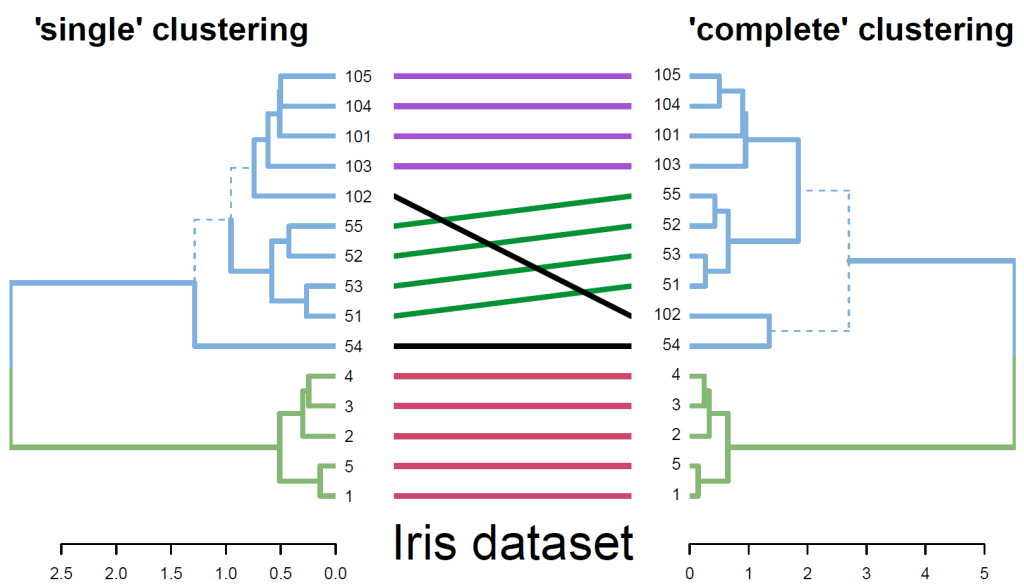
Fig. 2. A tanglegram for comparing two clustering algorithms used on 15 flowers from the Iris dataset. Similar sub-trees are connected by lines of the same color, while branches leading to distinct sub-trees are marked by a dashed line.
Comparing two dendrograms
The tanglegram function allows the visual comparison of two dendrograms, from different algorithms or experiments, by facing them one in front of the other and connecting their labels with lines. Distinct branches are marked with a dashed line. For easier and nicer plotting, dendlist concatenates the two dendrograms together, while untangle attempts to rotate trees with un-aligned labels in search for a good layout. Fig. 2 demonstrates a comparison of two clustering algorithms (single vs. complete linkage) on a subset of 15 flowers from the famous Iris dataset. The entanglement function measures the quality of the tanglegram layout. Measuring the correlation between tree topologies can be calculated using different measures with cor.dendlist (Sokal and Rohlf, 1962), Bk_plot (Fowlkes and Mallows, 1983), or dist.dendlist. Permutation test and bootstrap confidence intervals are available. The above methods offer sensitivity and replicability analysis for researchers who are interested in validating their hierarchical clustering results.
Enhancing other packages
The R ecosystem is abundant with functions that use dendrograms, and dendextend offers many functions for interacting and enhancing their visual display: The function rotate_DendSer (Hurley and Earle, 2013) rotates a dendrogram to optimize a visualization-based cost function. Other functions allow the highlighting of un-even creation of clusters with the dynamicTreeCut package (Langfelder et al., 2008), as well as of “significant” clusters based on the pvclust package (Suzuki and Shimodaira, 2006). Previously mentioned functions can be combined to create a highly customized (rotated, colorful, etc.) static heatmap using heatplot.2 from gplots (Warnes et al., 2014), or a D3 interactive heatmap using the d3heatmap package. The circlize_dendrogram function produces a simple circular tree layout, while more complex circular layouts can be achieved using the circlize package (Gu et al., 2014). Aside from R base graphics, a ggplot2 dendrogram may be created using the as.ggdend function.
In conclusion – the dendextend package simplifies the creation, comparison, and integration of dendrograms into fine-tuned (publication quality) graphs. A demonstration of the package on various datasets is available in the supplementary materials.
Acknowledgements
This work was made possible thanks to the code and kind support, of Yoav Benjamini, Gavin Simpson, Gregory Jefferis, Marco Gallotta, Johan Renaudie, The R Core Team, Martin Maechler, Kurt Hornik, Uwe Ligges, Andrej-Nikolai Spiess, Steve Horvath, Peter Langfelder, skullkey, Romain Francois, Dirk Eddelbuettel, Kevin Ushey, Mark Van Der Loo, and Andrie de Vries.
Funding: This work was supported in part by the European Research Council under EC–EP7 European Research Council grant PSARPS-297519. Conflict of Interest: none declared.
References
- Chipman,H. and Tibshirani,R. (2006) Hybrid hierarchical clustering with applications to microarray data. Biostatistics, 7, 286–301.
- Eddelbuettel,D. and François,R. (2011) Rcpp: Seamless R and C++ Integration. J. Stat. Softw., 40, 1–18.
- Fowlkes,E.B. and Mallows,C.L. (1983) A Method for Comparing Two Hierarchical Clusterings. J. Am. Stat. Assoc., 78, 553 – 569.
- Gu,Z. et al. (2014) circlize implements and enhances circular visualization in R. Bioinformatics, 30, 1–2.
- Hornik,M.M. and P.R. and A.S. and M.H. and K. (2014) cluster: Cluster Analysis Basics and Extensions.
- Hurley,C.B. and Earle,D. (2013) DendSer: Dendrogram seriation: ordering for visualisation.
- Langfelder,P. et al. (2008) Defining clusters from a hierarchical cluster tree: the Dynamic Tree Cut package for R. Bioinformatics, 24, 719–20.
- Milton,B.S. and Wickham,H. (2014) magrittr: magrittr – a forward-pipe operator for R.
- Paradis,E. et al. (2004) APE: Analyses of Phylogenetics and Evolution in R language. Bioinformatics, 20, 289–290.
- R Core Team (2014) R: A Language and Environment for Statistical Computing.
- Schmidtlein,S. et al. (2010) A brute-force approach to vegetation classification. J. Veg. Sci., 21, 1162–1171.
- Sokal,R.R. and Rohlf,F.J. (1962) The comparison of dendrograms by objective methods. Taxon, 11, 33–40.
- Suzuki,R. and Shimodaira,H. (2006) Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics, 22, 1540–2.
- Warnes,G.R. et al. (2014) gplots: Various R programming tools for plotting data.
- Wickham,H. (2011) testthat: Get started with testing. R J., 3, 5–10.
- Witten,D.M. and Tibshirani,R. (2010) A framework for feature selection in clustering. J. Am. Stat. Assoc., 105, 713–726.
no supplemental material found in the bioinformatics website.
Hi ygc,
You are right, they did not put it online yet.
I mentioned ggtree their briefly, but did not go deeper into it (or any of the other package for that matter).