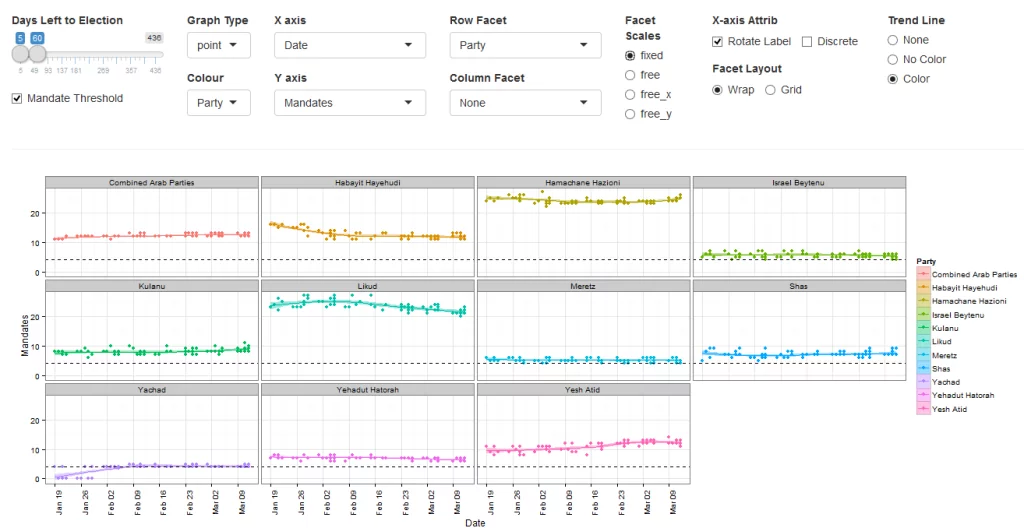
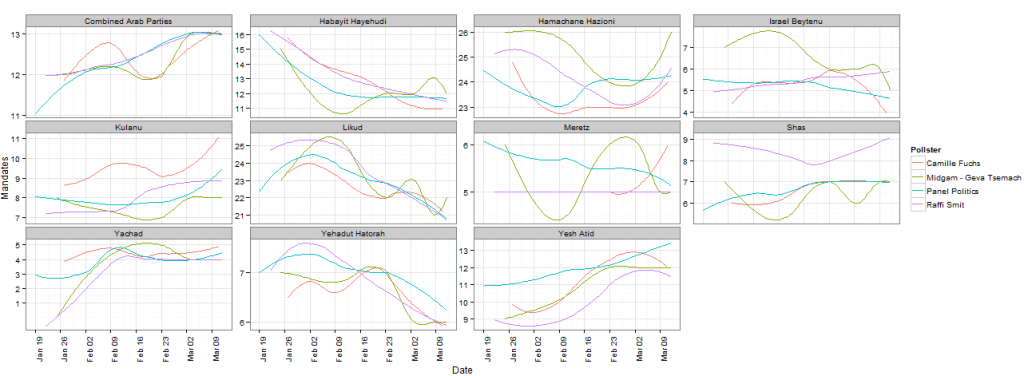
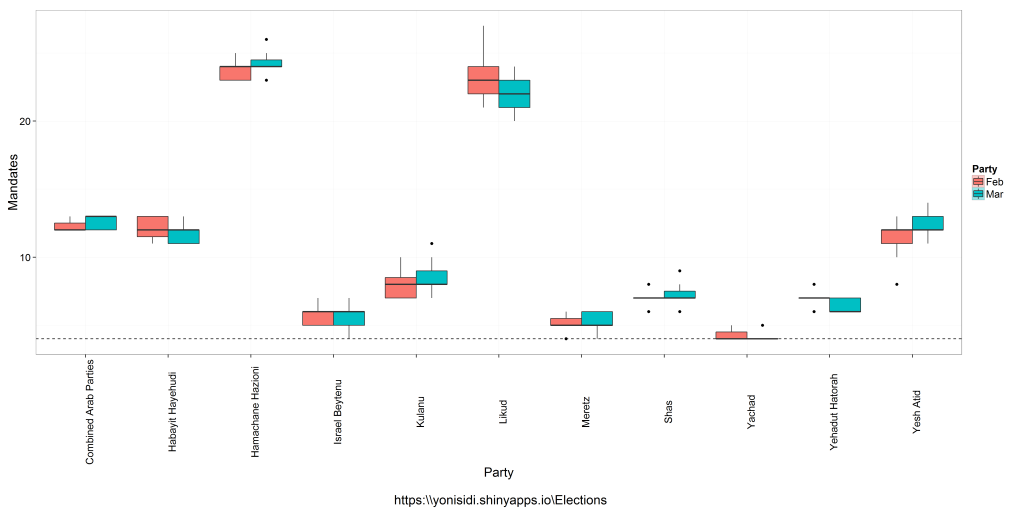
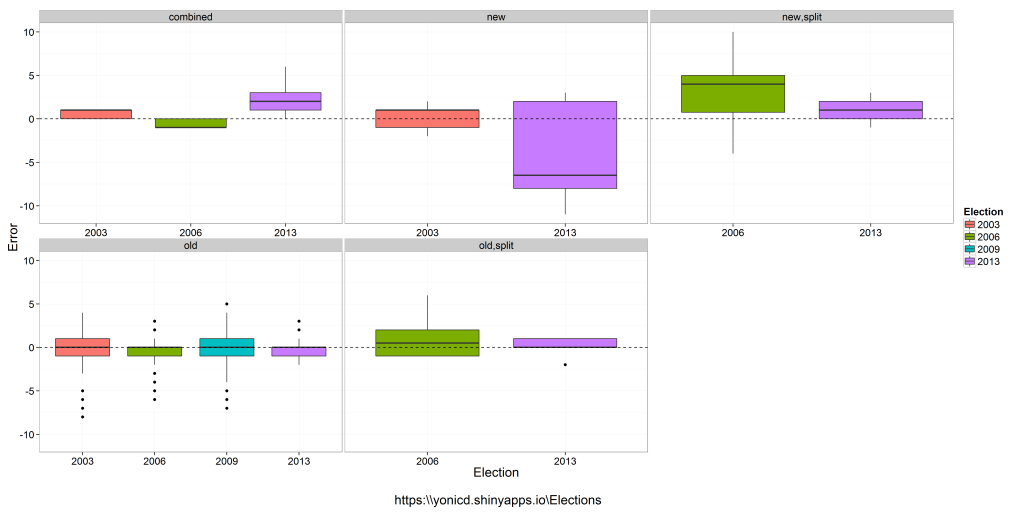
The followings introductory post is intended for new users of R. It deals with interactive visualization using R through the iplots package.
This is a guest article by Dr. Robert I. Kabacoff, the founder of (one of) the first online R tutorials websites: Quick-R. Kabacoff has recently published the book ”R in Action“, providing a detailed walk-through for the R language based on various examples for illustrating R’s features (data manipulation, statistical methods, graphics, and so on…). In previous guest posts by Kabacoff we introduced data.frame objects in R and dealt with the Aggregation and Restructuring of data (using base R functions and the reshape package).
For readers of this blog, there is a 38% discount off the “R in Action” book (as well as all other eBooks, pBooks and MEAPs at Manning publishing house), simply by using the code rblogg38 when reaching checkout.
Let us now talk about Interactive Graphics with the iplots Package:

Interactive Graphics with the iplots Package
The base installation of R provides limited interactivity with graphs. You can modify graphs by issuing additional program statements, but there’s little that you can do to modify them or gather new information from them using the mouse. However, there are contributed packages that greatly enhance your ability to interact with the graphs you create—playwith, latticist, iplots, and rggobi. In this article, we’ll focus on functions provided by the iplots package. Be sure to install it before first use.
While playwith and latticist allow you to interact with a single graph, the iplots package takes interaction in a different direction. This package provides interactive mosaic plots, bar plots, box plots, parallel plots, scatter plots, and histograms that can be linked together and color brushed. This means that you can select and identify observations using the mouse, and highlighting observations in one graph will automatically highlight the same observations in all other open graphs. You can also use the mouse to obtain information about graphic objects such as points, bars, lines, and box plots.
The iplots package is implemented through Java and the primary functions are listed in table 1.
Table 1 iplot functions
| |
| ibar() | Interactive bar chart |
| ibox() | Interactive box plot |
| ihist() | Interactive histogram |
| imap() | Interactive map |
| imosaic() | Interactive mosaic plot |
| ipcp() | Interactive parallel coordinates plot |
| iplot() | Interactive scatter plot |
To understand how iplots works, execute the code provided in listing 1.
Listing 1 iplots demonstration
1
2
3
4
5
6
7
8
9
10
11
12
| library(iplots)
attach(mtcars)
cylinders <- factor(cyl)
gears <- factor(gear)
transmission <- factor(am)
ihist(mpg)
ibar(gears)
iplot(mpg, wt)
ibox(mtcars[c("mpg", "wt", "qsec", "disp", "hp")])
ipcp(mtcars[c("mpg", "wt", "qsec", "disp", "hp")])
imosaic(transmission, cylinders)
detach(mtcars) |
Six windows containing graphs will open. Rearrange them on the desktop so that each is visible (each can be resized if necessary). A portion of the display is provided in figure 1.

Figure 1 An iplots demonstration created by listing 1. Only four of the six windows are displayed to save room. In these graphs, the user has clicked on the three-gear bar in the bar chart window.
Now try the following:
- Click on the three-gear bar in the Barchart (gears) window. The bar will turn red. In addition, all cars with three-gear engines will be highlighted in the other graph windows.
- Mouse down and drag to select a rectangular region of points in the Scatter plot (wt vs mpg) window. These points will be highlighted and the corresponding observations in every other graph window will also turn red.
- Hold down the Ctrl key and move the mouse pointer over a point, bar, box plot, or line in one of the graphs. Details about that object will appear in a pop-up window.
- Right-click on any object and note the options that are offered in the context menu. For example, you can right-click on the Boxplot (mpg) window and change the graph to a parallel coordinates plot (PCP).
- You can drag to select more than one object (point, bar, and so on) or use Shift-click to select noncontiguous objects. Try selecting both the three- and five-gear bars in the Barchart (gears) window.
The functions in the iplots package allow you to explore the variable distributions and relationships among variables in subgroups of observations that you select interactively. This can provide insights that would be difficult and time-consuming to obtain in other ways. For more information on the iplots package, visit the project website at http://rosuda.org/iplots/.
Summary
In this article, we explored one of the several packages for dynamically interacting with graphs, iplots. This package allows you to interact directly with data in graphs, leading to a greater intimacy with your data and expanded opportunities for developing insights.
This article first appeared as chapter 16.4.4 from the “R in action“ book, and is published with permission from Manning publishing house. Other books in this serious which you might be interested in are (see the beginning of this post for a discount code):